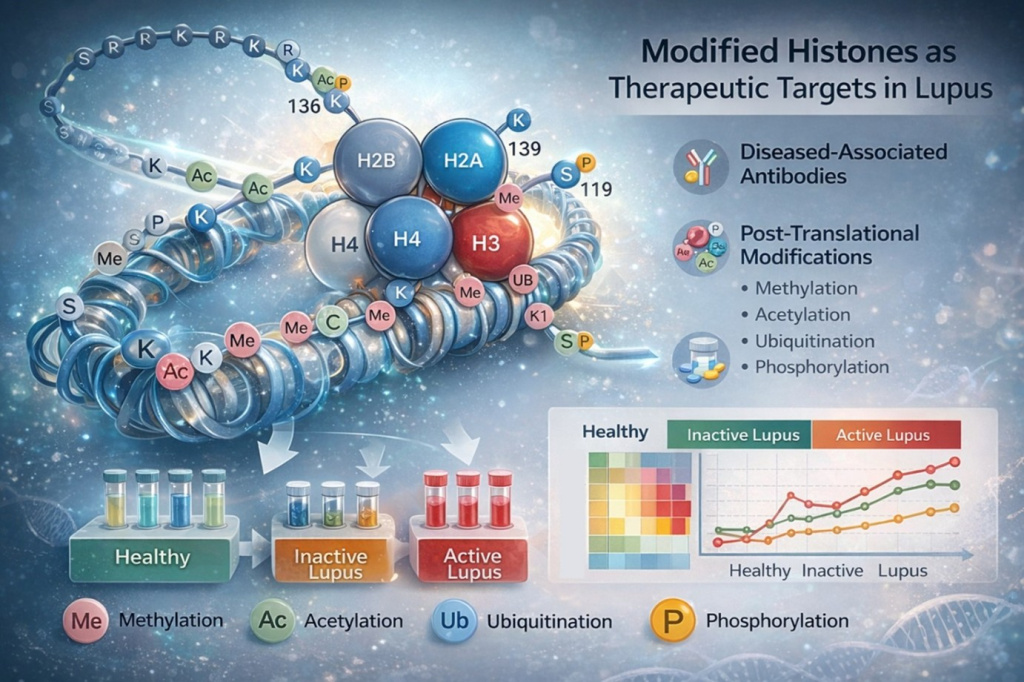
Post-Translationally Modified Histones as Therapeutic Targets in Lupus
Lead: Vinaika Maruvada
Team Members: Yewei Ma
Collaborators: Epicypher, Ramesh Saxena, Michelle Petri
Project Summary:
Systemic lupus erythematosus (SLE) is a complex autoimmune disease characterized by loss of immune tolerance and the generation of autoantibodies against nuclear antigens. Histones, which are core components of chromatin, are among the most well-recognized autoantigens in lupus. Importantly, histones undergo diverse post-translational modifications (PTMs), such as acetylation, methylation, and citrullination, which can alter chromatin structure, gene expression, and immunogenicity. This project investigates whether post-translationally modified histones represent disease-relevant immune targets in lupus and whether they may serve as diagnostic leads.
We analyze IgG and IgM antibody responses to histone and PTM-modified histone antigens using a multiplex Luminex platform. Samples include healthy controls, patients with inactive SLE, and patients with active lupus, enabling comparative analysis across disease states. By integrating and visualizing antibody reactivity patterns, this work seeks to identify histone modifications that are preferentially targeted in lupus, and/or associated with disease activity.
What is already known in the field?
- Autoantibodies targeting histones and chromatin components are a hallmark of lupus.
- Post-translational modifications can significantly influence histone structure, function, and antigenicity.
- Modified self-antigens are thought to contribute to loss of immune tolerance and sustained autoimmune responses, but their role in lupus remains unclear.
What is new?
- This project interrogates IgG and IgM antibody responses to a panel of histone and post-translationally modified histone antigens across multiple disease states.
- Combined heatmap visualizations and quantitative summaries are used to compare antibody reactivity patterns in healthy, inactive SLE, and active lupus groups.
- The study explores whether specific histone modifications are selectively recognized by the lupus immune response, suggesting biological relevance beyond unmodified histones.
Why is this important?
- Identifying disease-associated immune responses to modified histones may provide insight into mechanisms underlying lupus pathogenesis and disease heterogeneity.
- Histone modifications that show consistent, disease-linked antibody reactivity may serve as novel disease biomarkers for clinical use.
- Identifying the earliest PTM epitopes targeted in lupus may point to the mechanistic origins of the disease.
Ongoing/Future Steps
- Initial exploratory data analysis of combined IgG and IgM antibody responses has been completed, including data integration, visualization, and group-wise comparisons across disease states.
- Perform validation studies in additional patient samples, using multiplexed Luminex assays.
- Identify histone modifications that correlate with disease activity or immune activation, or identify the pre-diagnosis state.
- Evaluate the potential of selected modified histones as candidates for therapeutic targeting or point-of-care biomarker assays.